Speed Up Sparse Boolean Data
05 Jan 2023Table of Contents
I’m working on replicating the (Re-)Imag(in)ing Price Trends paper - the idea is to train a Convolutional Neutral Network (CNN) "trader" to predict the stocks' return. What makes this paper interesting is the model uses images of the pricing data, not in the traditional time-series format. It takes financial charts like the one below and tries to mimic the traders' behaviours to buy and sell stocks to optimise future returns.
Alphabet 5-days Bar Chart Shows OHLC Price and Volume Data
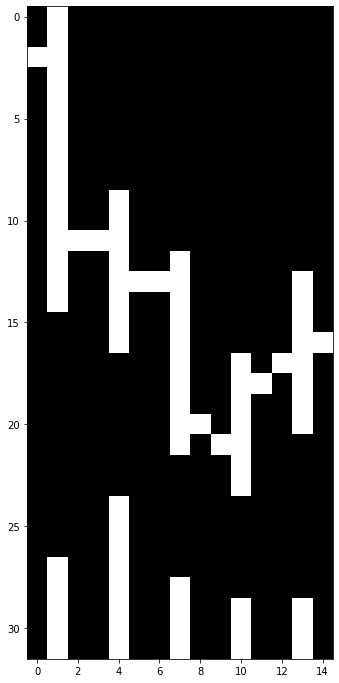
I like this idea. So it becomes my final assignment for Deep Learning Systems: Algorithm and Implementations course.
Imaging On-the-fly
To train the model, the price and volume data are transformed into black-white images which is just a 2D matrix with 0s and 1s. For just around 100 stocks' pricing history, there are around 1.2 million images in total.
I used the on-the-fly imaging process during training: in each batch, it loads pricing data for a given stock, sample one day in the history, slice a chunk of pricing data, and then convert it to an image. It takes about 0.2 milliseconds (ms) to do all that, so in total it takes 4 minutes to loop through all the 1.2 million images.
%%timeit
df = MarketData(DATA_DIR)['GOOGL']
imager = ImagingOHLCV(img_resolution, price_prop=price_prop)
img = imager(df.tail(5))
1.92 ms ± 26.9 µs per loop (mean ± std. dev. of 7 runs, 1,000 loops each)
To train 10 epochs, that's 40 minutes in loading data. To train one epoch on the full dataset with 5,000 stocks, that's 200 minutes in loading data alone!
PyToch utilises multiple processing in loading the data using CPU while training using GPU. So the problem is less severe, but I'm using the needle, the deep learning framework we developed during the course, it does have this functionality yet.
During training using needle, the GPU utilisation is only around 50%. After all the components in the end-to-end are almost completed, it is time to train with more data, go deeper (larger/more complicated morel), try hyper-parameters tuning etc.
But before moving to the next stage, I need to improve the IO.
Scipy Sparse Matrix
In the image above, there are a lot of black pixels or zeros in the data matrix. In general only 5%-10% of pixels are white in this dataset.
So my first attempt was to use scipy's spare matrix instead of numpy's dense matrix: I save the sparse matrix, loaded it, and then convert it back to a dense matrix for training CNN model.
%%timeit
img_sparse = sparse.csr_matrix(img)
sparse.save_npz('/tmp/sparse_matrix.npz', img_sparse)
img_sparse_2 = sparse.load_npz('/tmp/sparse_matrix.npz')
assert np.all(img_sparse_2 == img)
967 µs ± 4.99 µs per loop (mean ± std. dev. of 7 runs, 1,000 loops each)
It reduces the IO time to 1ms, so about half of the time, not bad, but I was expecting a lot more as the sparseness is high.
Numpy Bites
Then I realised the data behind images is just 0 and 1, in fact, a lot of zeros, and only some are 1. So I can ignore the 0s and only need to save those 1s, then reconstruct the images using those 1.
It is so simple that numpy has functions for this type of data processing already. The numpy.packbites function converts the image matrix of 0 and 1 into a 1D array whose values indicate where the 1s are. Then the numpy.unpackbits does the inverse: it reconstructs the image matrix by using the 1D location array.
This process reduces the time of loading one image to 0.2 milliseconds, that's 10 times faster than the on-the-fly method with only a few lines of code.
%%timeit
temp_file = "/tmp/img_np_bites.npy"
img_np_bites = np.packbits(img.astype(np.uint8))
np.save(temp_file, img_np_bites)
img_np_bites = np.load(temp_file)
img_np_bites = np.unpackbits(img_np_bites).reshape(img.shape)
assert np.all(img_np_bites == img)
194 µs ± 3.95 µs per loop (mean ± std. dev. of 7 runs, 10,000 loops each)
Another benefit is the file size is much smaller: it is 188 bytes compared to 1104 bytes using sparse matrix. So it takes only 226MB of disk space to save 1.2 million images!
Path('/tmp/img_np_bites.npy').stat().st_size, Path('/tmp/sparse_matrix.npz').stat().st_size
188, 1104
Problems of Having Millions of Files
It takes a couple of minutes to generate 1.2 million files on my Debian machine. It is so quick! But then I release this approach is not scalable without modification because there's a limited number of files the OS can accommodate. The technical term is Inode. According to this StackExchange question, once the filesystem is created, one cannot increase the limit (Yes, I was there).
Without going down to the database route, one quick workaround is to bundle the images together, for example, 256 images in one file. So later in training, load 256 images in one go, then split them into chunks. Just ensure the number of images is a multiple of the batch size used in training so I don't have to deal with unequal batch sizes. Since those bundled images are trained together, it reduces the randomness of SGD, so I won't bundle too many images together, 256 sounds about right.
The LSP and other tools can cause problems when they are monitoring folders with a large number of files. Moving them out of the project folder is the way to go so Emacs won't complain or freeze.